Institut Cochin :
Equipe Développement neuromusculaire, génétique et physiopathologie
https://www.institutcochin.fr/la-recherche/drc/equipe-maire-sotiropoulos
https://www.institutcochin.fr/la-recherche/drc/equipe-maire-sotiropoulos
Pascal Maire (DR1 INSERM),
Athanassia Sotiropoulos (DR2 INSERM)
Evelyne Bloch-Gallego (DR2 INSERM), Florian Britto (MCU, UP), Thierry Launay (MCU, UP), Stéphanie Backer (IEHC, INSERM), Nathalie Chaverot (AI, CNRS), Iryna Pirozhkova (IR, CNRS en mission), Voahangy Randrianarison (IRHC, CNRS)
Laura Ben Driss (Doctorante, UP), Léa Délivry (Doctorante, UP), Kassandra Kobon (Doctorante, UP), Rouba Madani (Doctorante, UP)
Muscle, exercice, développement, exercice, hypertrophie, atrophie
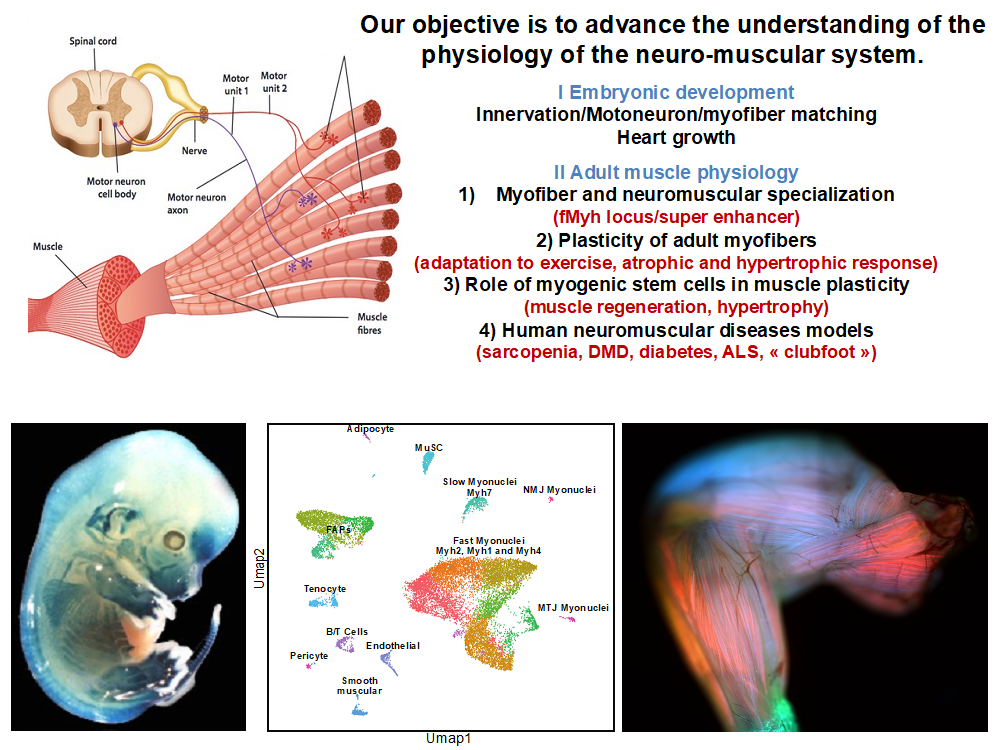
The adult skeletal muscle tissue is composed primarily of specialized cells, myofibers, which represent 40-50 % of body weight in humans. They have associated muscle stem cells (MSC) called satellite cells. The muscles are innervated by motoneurons, which synaptic activities lead to muscle contractions and motor activity. In Europe, more than 15 million people are affected by muscle diseases that always evolve to bring a loss or atrophy of muscle tissue. These diseases are either genetic or accompany hospitalization or prolonged immobilization or are the result of various diseases. In addition, the increase of life expectancy during the last century has caused an augmentation of pathologies emergence including cancer, obesity, type 2 diabetes, neurodegenerative and cardiovascular diseases. Most of them are associated with alteration of muscle function and/or sarcopenia. The alteration of muscle function is consecutive to myofibers loss or atrophy. Importantly, physical exercise improves muscle function and locomotion, and physical training is now considered as one of the most effective non-pharmaceutical strategy to slow down muscle wasting. Various diseases also affect the development and survival of neurons. This is particularly the case of amyotrophic lateral sclerosis (ALS) and spinal muscular atrophy characterized by muscle weakness.
Projects developed within our team aim to characterize the influence of transcription factors, guidance factors and signaling pathways in muscle fibers and in their environment (including MSC and motoneurons) during development and in the adult in response to loss or gain in activity to understand their implications in normal and pathological development of skeletal muscle, and in motoneurons. These studies are based on the use of models of mutant mice for genes of interest, or in cellular models. We hope to get an integrated picture of the response patterns of the neuromuscular system to its environment under normal and pathological conditions through among others high throughput genomic studies (snRNA-seq, snATAC-seq).
[1] Dos Santos,M., Backer,S., Saintpierre,B., Izac,B., Andrieu,M., Letourneur,F., Relaix,F., Sotiropoulos,A., Maire,P. 2020. Single-nucleus RNA-seq and FISH identify coordinated transcriptional activity in mammalian myofibers. Nat Commun 11, 5102 (2020).
[2] Wurmser,M., Chaverot,N., Madani,R , Sakaï,H., Negroni,E., Demignon,J., Saint-Pierre,B., Mouly,V., Amthor,H., Tapscott,S., Birchmeier,C., Tajbakhsh,S., Le Grand,F., Sotiropoulos,A., Maire,P. 2020. SIX1 and SIX4 homeoproteins regulate PAX7+ progenitor cell properties during fetal epaxial myogenesis. 147(19):dev185975. doi: 10.1242/dev.185975.
[3] Montel L, Sotiropoulos A, Hénon S. 2019. The nature and intensity of mechanical stimulation drive different dynamics of MRTF-A nuclear redistribution after actin remodeling in myoblasts. PLoS One. 14(3):e0214385. doi: 10.1371/journal.pone.0214385. eCollection 2019.
[4] Backer S, Lokmane L, Landragin C, Deck M, Garel S, Bloch-Gallego E. 2018. Trio GEF mediates RhoA activation downstream of Slit2 and coordinates telencephalic wiring. Development. 145(19):dev153692. doi: 10.1242/dev.153692.
[5] Randrianarison-Huetz, V. *, Aikaterini Papaefthymiou, A. *, Herledan, G., Noviello, C., Faradova,U., Collard, L., Pincini,A., Schol,E., Decaux, J-F., Maire, P., Vassilopoulos, S., Sotiropoulos, A. 2018. Srf controls satellite cell fusion through the maintenance of actin architecture. J.Cell.Biol. 217(2):685-700